
Adrian's Rojak Pot has long being a participating team in the Distributed Folding Project. Our Team started off when the Project was in its infancy and we have continually supported it even though we are only a small team. We hope that with our contribution, the Project will be able to make significant contributions to the development of future drugs that can help cure many of today's diseases.
Now, this is not a classified research project limited to a select group of people. Believe it or not, you and everyone else can be a part of this visionary project! All you need is your computer, an Internet connection and a few minutes of your time. Read on to find out more!
What is Distributed Folding?
Short Answer: Distributed Folding is a research project that uses your computer's unused clock cycles to help research protein folding. Scientists need to know how they fold so that they can find ways to create better drugs.
Long Answer: Distributed Folding is a project that uses the distributed computing concept (instead of a supercomputer) to reduce the time and cost required to solve a monumental problem. It does so by distributing the task or job to a multitude of machines, all connected via the Internet! This allows the Distributed Folding Project team to study the many ways a protein can fold without the need for a supercomputer! You can read more about the wonders of protein folding at the Distributed Folding website!
How do I run Distributed Folding?
Running Distributed Folding (or as it is affectionately know as - DF) is very simple. All you need is a computer with these minimum system requirements :-
- 233 MHz processor or faster (for Windows versions, at least a Pentium II processor or equivalent is recommended)
- 20 MB free hard drive space (up to 100 MB additional temporary space)
- 128 MB RAM
- An Internet connection (broadband or 14.4k or faster modem)
- A supported operating system (Windows 95/98/ME/NT/2000/XP, Linux, FreeBSD, Solaris, IRIX, Macintosh and many others)
If you satisfy the minimum requirements above, you are all set and ready to go!
You will need the DF client which can be downloaded at the Distributed Folding website. Choose the protein client that supports your operating system. If you have a Microsoft Windows machine, all you need to do is click on this link to download the client directly from the official Distributed Folding FTP (File Transfer Protocol) site.
You will then have to register yourself with the Distributed Folding server before you can start to fold proteins. The registration process is a simple one where you are required to fill in an e-mail address, a password and the organization you are affliated with. You can register yourself here!
After registration, you will be given an 8-character code, called the handle. Remember your handle as it is unique to you and it will ensure that the DF server recognizes and authenticates you. If you forget, that's alright as you can request that your handle be sent to your e-mail address.
Selecting The Distributed Folding Client
If you are using a Microsoft Windows operating system, you will have two types of DF clients to choose from. One is in the form of a screensaver while the other is a text client.
If you are looking to get maximum output, it is recommended that you get the text client as it processes proteins faster. This is because it does not need to render lovely representations of your proteins!
But if you would like to use it as a screensaver and see your proteins folding on your screen, then feel free to get the screensaver client instead!
Assuming that you have chosen to use the text client, you will have a ZIP file called "distribfold-current-win9x.zip". It should be around 8MB in size.
Unzip the contents of this file to a location that you can remember. It is recommended that you unzip it to the desktop if you are unsure of how to do it.
Once you are done, you will see a folder called "distribfold". Double-click on the folder and you will see the files for the Distributed Folding client.
Find the "foldit.bat" file and double-click it to start up the client. The client will start in a DOS window with a black screen. It will then ask you for your handle.
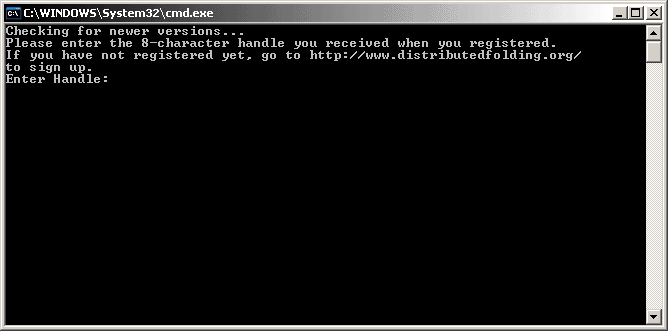
The handle is the 8-character code provided by the Distributed Folding server after you have registered. Type it and press Enter.
The Distributed Folding client will authenticate and register it, thereby allowing you to receive credit for proteins folded by that client.
Configuring the Distributed Folding Client
After authenticating yourself, you will be asked a series of questions to configure the DF client.
First is the Quiet Mode option.
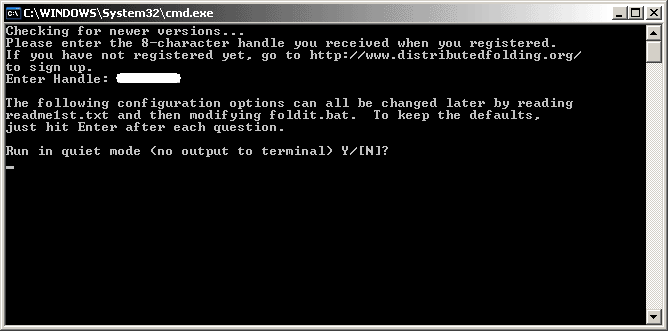
This mode allows the client to run without displaying the ASCII protein "snake" in the DOS box. If you are a first time user, it will be good to run without the quiet mode to check out the output. You will be impressed with the display. So, press "N" and then Enter.
Next, the DF client will ask you if you want it to Connect to the Internet automatically to upload proteins.
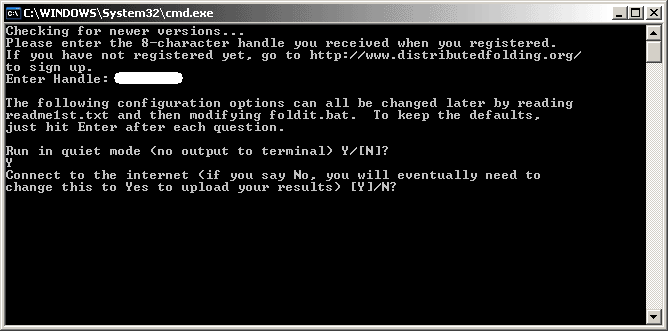
If you are a dialup user, you will want to type "N" to prevent the DF client from automatically connecting to the Distributed Folding server to upload its proteins. However, you must remember to connect yourself to the Internet when you want to upload proteins that the client has folded. Press "N" and Enter.
If you are connected to the Internet all the time (i.e. via DSL or cable), type"Y" and Enter to allow the DF client to automatically connect to the server and upload its proteins.
The next option will determine if the DF client is allowed to use extra memory of up to 150MB. This option allows the client to be almost twice as fast at folding proteins, provided you have enough memory in your computer, of course!
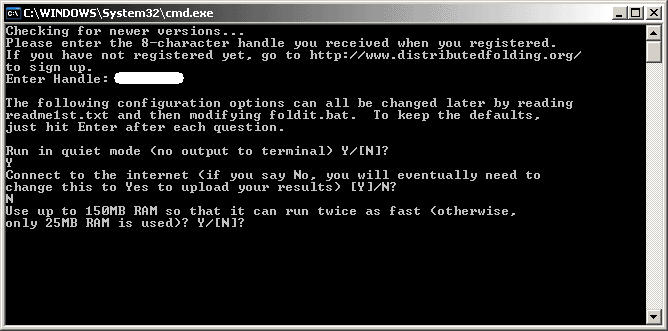
For most users, it is recommended that you select "N" for this option. The DF client may not be as fast at folding proteins but it will use very little memory.
But if you are all out to help fold proteins for the Project, you should select "Y"! We highly recommend that you only enable this option if you have at least 512MB of memory.
Next up is the option to automatically accept updates from the DF server.
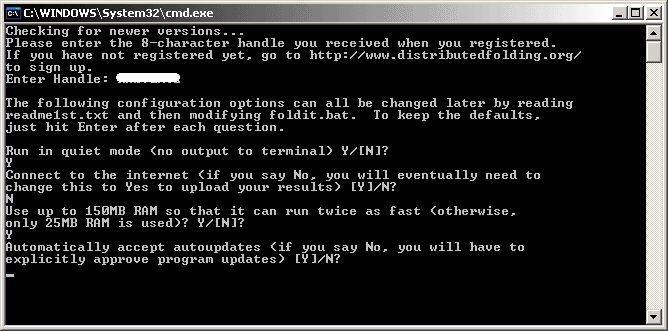
If select "Y", the client will auto-update itself when a new update is detected. All updates are digitally signed by the Distributed Folding Team so it's safe to accept auto-updates.
It is highly recommended that you select "Y".
The final option allows you to configure a HTTP Proxy server.
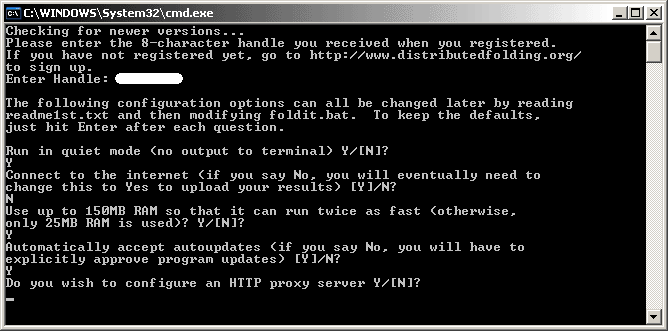
If you have no idea what that means, chances are that you don't need this feature! Press "N" and Enter.
The DF client will now announce that "Setup is complete"!
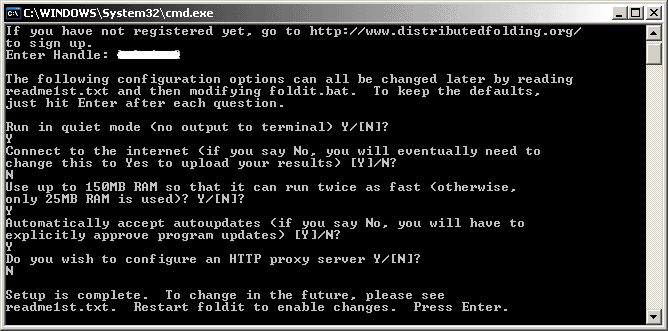
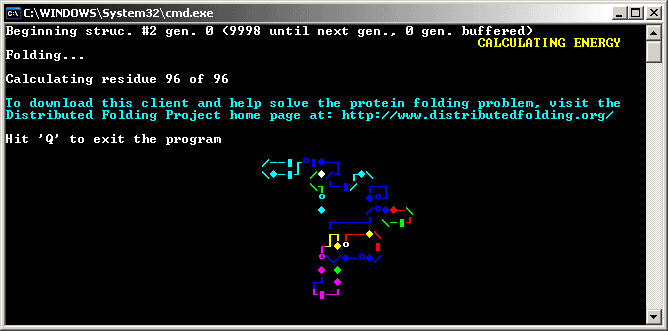
Press Enter to close the client and then double-click the "foldit.bat" file again to restart the client.
If you did not enable the "Quiet Mode", you will see an "ASCII protein snake" in the DOS box. Congratulations! You have just taken the first step in helping mankind better understand how proteins fold!
dfGUI
We highly recommend that you download dfGUI and install it.
It is a helper application for Windows and Linux that allows you to easily change the client's settings. You can also use it to AutoStart, AutoStop and Restart the client in the event of a crash. It will also Recover the client if the the client was not shutdown properly. dfGUI is also able to to hide in the Tasktray and hide the DOS box of the client.
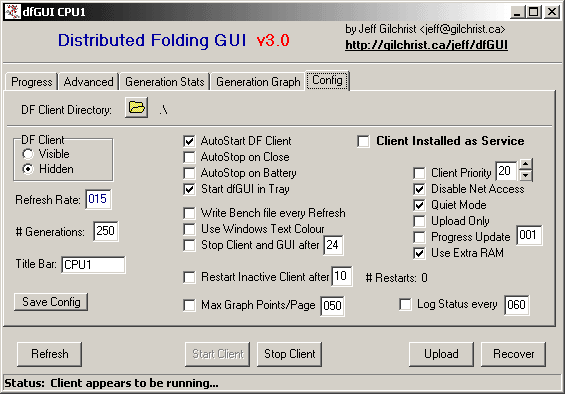
We find it extremely helpful in configuring and controlling the client. Many thanks to Jeff Gilchrist for this wonderful application! If you have problem using dfGUI, please visit the official Distributed Folding forum where Jeff is at hand to answer your questions.
More Information
You can obtain more information from the readme1st.txt file in the distribfold folder or at the Distributed Folding website. The FAQ for the Phase II client can be found here and if you are still lost or need help, please visit the official forum for the Distributed Folding Project here.
As always, we are here to answer any questions you might have. Our Team ARP Distributed Folding forum is here. If you ever need a question answered or if you would like to join our team, please do contact us here!.
Although our team is comparatively small, we have great fun trying to outdo each other in folding the most proteins! Try it and you will see what we mean! But whatever your reason for joining this noble cause, we all thank you! :)






 Add to Reddit
Add to Reddit
